 The first step in gating is often distinguishing populations of cells based on their forward and side scatter properties. In addition to this flow guide we have a handy PDF available to download on our gates, plots and regions page, which can be used as a quick reminder. Here we will show what the common flow cytometry graph outputs look like and how in a few simple steps you can identify different cell populations that have been stained with antibodies conjugated to fluorophores. Gates and regions are placed around populations of cells with common characteristics, usually forward scatter, side scatter and marker expression, to investigate and to quantify these populations of interest. If a researcher is interested in how age affects CD28 expression, change “parameter” argument from “fraction” to “CD28”.Flow cytometry data analysis is fundamentally based upon the principle of gating. In this case, we are interested in how age affects the fraction of cells. the “parameter” variable specifies the type of summary statistics. the study ID variable will always be included to adjust for batch effects. In this case, Gender is included to control for gender differences. The “otherVariables” argument specifies other variables to be included in the regression. The function will only report the effect size for the “variableOfInterst”, which is Subject Age in this case. The “variableOfInterest” argument specifies the variable that a researcher is interested in. The following code performs a regression model: fraction ~ Subject Age + Gender + study_id. To see how the fraction of cell subsets within the blood chance with age, we can use the glmAnalysis function. You won't need this line when running your actual meta-analysis sample_info $fcs_files= system.file( "extdata",sample_info $fcs_files, package= "MetaCyto") # join the cluster summary statistics with sample information all_data= inner_join(fcs_stats,sample_info, by= "fcs_files") Library(dplyr) # Collect Summary statistics generated in step 3 files= list.files( "Example_Result/search_output", pattern= "cluster_stats_in_each_sample", recursive= TRUE, full.names= TRUE) fcs_stats= collectData(files, longform= TRUE) # Get sample information generated in step 1 fn= system.file( "extdata", "sample_info_vignette.csv", package= "MetaCyto") sample_info= read.csv(fn, stringsAsFactors= FALSE, check.names= FALSE) # find data in the MetaCyto package. The path of fcs files in fcs_info.csv and sample_info.csv must be the same. So we will include gender and age in the file. In this example, we try to see how gender and age affect immune cells. The second file (sample_info.csv) contains the information about the samples or subjects corresponding to each of the fcs files. SDY420/ResultFiles/CyTOF_result/11462_cells_ 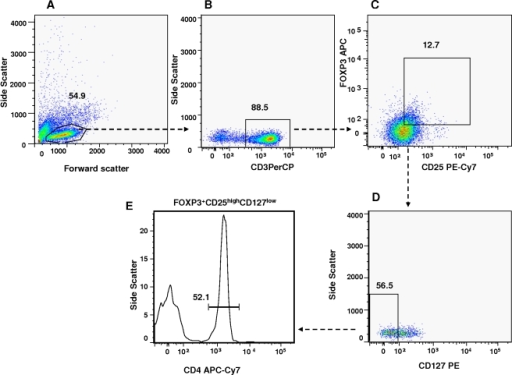
SDY420/ResultFiles/CyTOF_result/11445_cells_ SDY420/ResultFiles/CyTOF_result/11059_cells_ The first file (fcs_info.csv) contains the path (relative to working direction) of each fcs files and the study each fcs file belongs to. The rest of steps in performing meta-analysis on ImmPort data is exactly the same as Example 1.įor your local datasets, the data collection has to be done manually (If you are analyzing data from ImmPort database, this step is automated, see example 2). This example shows how to automate the data collection step for cytometry data downloaded from ImmPort. This example shows how to perform meta-analysis using MetaCyto on your local datasets.Įxample 2: ImmPort Cytometry Datasets. More self-contained examples, including data and code, are available on GitHub (/hzc363/MetaCyto_Examples).Įxample 1: Local Cytometry Datasets. In the second example, we provide an example to show how to use functions in MetaCyto to automate the data collection step for datasets from ImmPort. The example will walk you through all 4 steps of meta-analysis. 
We created 2 examples to demonstrate the use of MetaCyto in this Vignette.In the first example, we provided an example to show how MetaCyto can be used to analyze your local cytometry datasets. MetaCyto carries out the meta-analysis in 4 steps: data collection, data pre-processing, identifying common cell subsets across studies and statistical analysis. It is able to jointly analyze cytometry data from different studies with diverse sets of markers. MetaCyto is an R package that performs meta-analysis of both flow cytometry and mass cytometry (CyTOF) data. 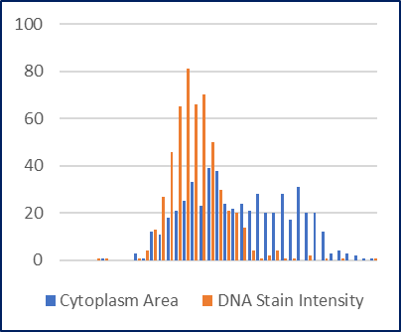
Introduction to MetaCyto Introduction to MetaCyto Zicheng Hu 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
